Difference between revisions of "The Biolibrarian Proposal"
| Line 1: | Line 1: | ||
| + | [[Image:ancientlibraryalex.jpg|right|Alexander's library]] | ||
| + | |||
== The survey == | == The survey == | ||
Revision as of 20:01, 6 April 2009
Contents
- 1 The survey
- 2 The Biolibrarian
- 3 What is the Biolibrarian proposal?
- 4 The Biolibrarian's role defined
- 5 How do Biolibrarians differ from Biocurators and Bioinformaticians?
- 6 Guiding principals of the Biolibrarian proposal
- 7 Existing sociological problems addressed by the Biolibrarian proposal
- 8 The proposal in brief
- 9 How to add your university to this survey
- 10 About the author and contact information
- 11 Related sites
The survey
Please take the Biolibrarian proposal survey by visiting http://www.surveymonkey.com/s.aspx?sm=LubRkhzbX9a7e8aPGKGevQ_3d_3d
It's only six questions and can be filled out in less than five minutes.
The Biolibrarian
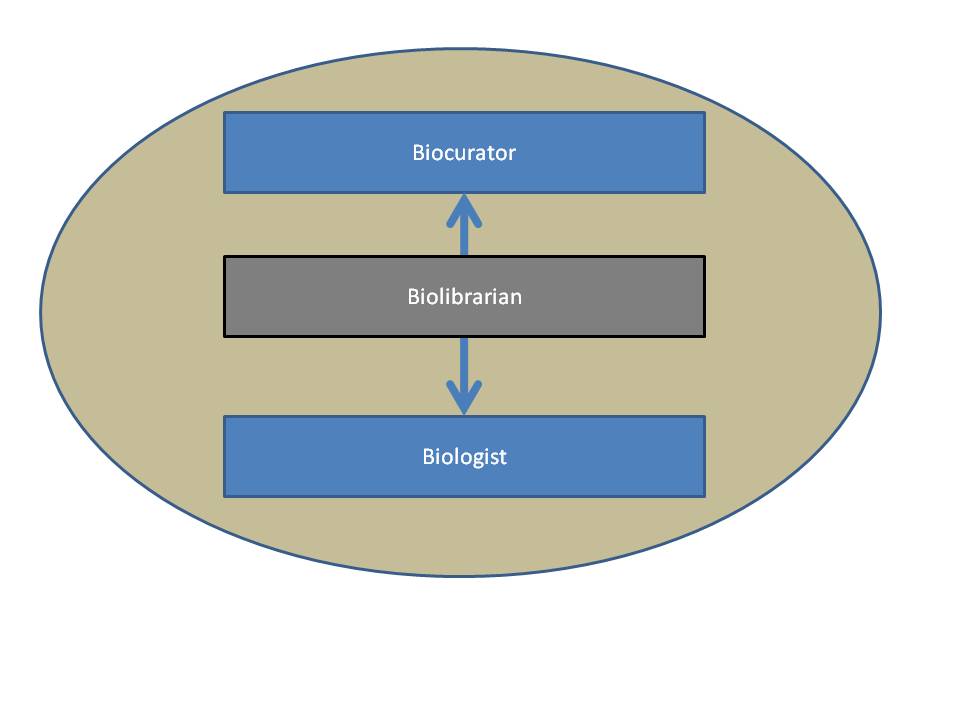
The Biolibrarian takes ownership for the transfer of information between biologists and biological databases.
What is the Biolibrarian proposal?
The Biolibrarian is a proposed new infrastructure position at university libraries around the world.
A Biolibrarian is trained in the use of biological databases and acts to
1) help biologists locate and use biological databases
2) help biologists submit data and feedback to biological databases
It is envisioned that molecular biologists could meet with a Biolibrarian in the same way that they meet with and use the services of a librarian. For example, the Biolibrarian could help a molecular biologist researcher to locate pathways, complexes and interactions that their molecules of interest are involved in.
In addition, the Biolibrarian could help biologists users to locate and submit data and feedback to biological databases.
For example, the Biolibrarian would be trained in the use of state of the art text mining tools to help researchers locate data for their molecules of interest in abstracts and full-text research articles. They could then help researchers enter verified information from full-text articles into curated databases where it would be available to researchers around the world that were querying for information on these same molecules.
In this role, the Biolibrarian would act as a broker between local researchers and database curators. This would apply to those databases that are set up to accept feedback and entries from external sources. Biolibrarians would also become local brokers that provide feedback on databases, interfaces and associated search tools.
Would you support such a service at your local university library? Do you have comments on this proposal? Follow the link above and take our survey.
You can read a synopsis of the proposal below.
Comments and suggestions are welcome. We are also interested in learning about similar proposals or projects that are already in place. Please email ian.donaldson@biotek.uio.no. If you would like to add your name to this wiki page in support of this application, please send a brief email.
The Biolibrarian's role defined
Biolibrarians:
act as brokers between biologists and databases: this means that they facilitate interaction between local biologists and biological databases. This is a two-way process that includes helping bioloigists find and use biological databases AND helping biologists submit data and feedback to biological databases.
facilitate retrieval of biological information: biolibrarians would be trained with specialized knowledge of major biological databases, that includes their data representations, data exchange mechansisms with other databases, and the controlled vocabularies and ontologies that they employ. These are all important aspects of being able to find, use and interpret biological data. Examples of major biological databases include databases like GenBank, UniProt, the Saccharomyces Genome Database, Genetic Association Database, Gene Ontology Database, The Protein Databank, The Eukaryotic Linear Motif Database, GAPScreener, IntAct, BioModels Database and Reactome. These are just a few of the hundreds of biological databases that may be of relevance to local biologists (see NAR online molecular biology database collection and PMID 19033364).
facilitate submission of data to biological databases: Each of the databases listed above will accept record submissions and/or feedback from biologist users. The quality of these databases is dependent on this relationship and many of these databases face critical shortages in curation personnel. The Biolibrarian's role is to to actively seek out and engage biologists in submitting data and feedback to these databases. The sheer number of databases, data models, and controlled vocabularies employed by these databases requires a specialist intermediary to facilitate this exchange of information.
How do Biolibrarians differ from Biocurators and Bioinformaticians?
Biologists, BioLibrarians and Biocurators are all Bioinformaticians. Each group both consumes and produces biological information. Biologists are the primary producers of biological information. Biocurators care for some specific subset of biological information associated with a specific database. Biolibrarians care for the transfer of knowledge between biologists and databases (Biocurators) regardless of the focus of the biologist or the database. Bioinformatics encompasses all aspects of this process of data production and consumption. The defining and differentiating aspects of the Biolibrarian are:
1) BioLibrarians are non-partisan with respect to databases (in contrast to biocurators): this means that biolibrarians will choose databases to submit data to based on their their appropriateness to the data at hand to be archived. These databases may include, for example, protein databases such as UniProt, EntrezGene or the Gene Ontology for functional annotation of gene products or the FACCS database for cataloging of the structure and structure-function relationships in the cerebro-cerebellar system. Biolibrarians may review and recommend databases and be involved in discussions around the use of and changes to database representations and controlled vocabularies.
2) Biolibrarians do not write code and are non-partisan with respect to development groups (in contrast to bioinformatician researchers). This means that Biolibrarians are primarily users of databases and associated software. They may review existing software or even tender the creation of new software either locally or internationally . Here, their knowledge of domain representation and query use-cases in a specific area would be used to shape requirements. However, Biolibrarians would not use or endorse software solely because it is produced and maintained by a research group at their local university.
3) Biolibrarians facilitate submission of data from biologists to databases (in contrast to bioinformatcian support groups). This is an active (not a passive) mandate. Biolibrarians would actively seek out and engage local researchers to collect and curate data. These data may be of general biological interest or specific to the local research group or to the specialization of the local Biolibrarian group. Again, the Biolibrarian acts as part of an infrastructure group that is local to the university but independent of any given research group.
Biolibrarians may already exist as part of bioinformatics service groups if they facilitate both data use and submission without regard to a specific data type or database.
Guiding principals of the Biolibrarian proposal
The following "guiding principals are all seen as essential to the development and sustainability of biolibrarian activities.
Biolibrarians are:
Local: they are part of research infrastructure at universities.
Global: biolibrarian groups may exist at any university with a biological research programme.
Non-partisan: biolibrarians do not represent any specific biological database or software development group. They are independent users and supporters of these groups.
Pro open source: where possible and appropriate, biolibrarian groups would favour use of open source software. This is meant to provide support for these projects and help ensure their continued availability to users.
Pro open access: where possible and appropriate, biolibrarian groups would favour use of open source databases and making their own work open access.
Pro standards: where possible, biolibrarian groups would adhere to established practices and data model usage. They may also act to establish or improve existing standards for data representation in cooperation with biological databases and their users.
Pro documentation: where possible, biolibrarian groups will collect and provide references to or training in documentation of standard curation practices for a data type. They may also act to establish or improve existing documentation in cooperation with biological databases and their users.
Active: biolibrarians would be mandated to actively seek out and engage researchers in the archiving and curation of biological data.
Authors: curation of biological data is a legitimate academic activity. Biolibrarians should be encouraged to author and co-authors papers with other researchers on their activities.
Existing sociological problems addressed by the Biolibrarian proposal
Ownership of biological data tranfer: Some biological data types (such as DNA sequence data) cannot be published as part of a research paper without first depositing the data in a recognized database. This community policy is enforced by journal editors. However, the overwhelming majority of biological data types do not fall under this community policy. We cannot expect that journals can or should enforce deposition of biological data into databases prior to publication; there are many databases that accept deposition but few of them have attained the historical long-term support and recognition shared by sequence databases such as GenBank, EMBL and DDBJ. This proposal argues that universities must be pro-active in the deposition of biological data; they are the producers of these data and hold the responsibility, the means and the incentive that best ensures that these data are captured in formats where they can be readily accessed by human researchers and machine algorithms.
Overwhelmingly large number of biological databases: The present scientific granting structure allows for a plethora of databases that range in complexity from proof-of-principal, protoypical databases generated as part of a M.Sc. project to well-established, permanently funded, multi-national consortiums. The typical biologist researcher cannot be expected to distinguish between or even be aware of these myriad efforts. This proposal argues that a specialized force of curators deployed at universities world-wide would be best suited to reviewing these databases and supporting data transfer to the most appropriate and well-established efforts.
Absence of stable funding for biological databases curation: Many excellent database efforts exist that lack stable and sufficient funding or even appropriate awareness from their respective potential user communities. This situation exists largely due to the fact that database projects often compete in the same space as experimentalist projects and fail to gain adequate funding on the grounds that they are "mere" infrastructure projects (despite the fact that these same databases are often used and cited by experimentalist research projects). This proposal argues that these databases, even under the best funding circumstances, cannot hope to cope with the required curation load in their respective areas. We propose that the best of these efforts deserve international support from curators that are local to universities around the world. These curators (or biolibrarians) would act to engage and direct attention of biological researchers to those database efforts that are most deserving of support. This same group of local biolibrarians would act to accelerate development of these efforts by providing multiple, real-world, use-case scenarios.
The proposal in brief
Time plan: 5 years
Personnel: 8 – 10 biolibrarian curators
Deliverable: A prototype for biolibrarian positions at universities around the world (with a special empahsis of protein interaction and biological pathway curation).
We will propose a team of 8 to 10 curators who will search primary biomedical research literature and enter biomolecular interaction and pathway data into machine readable format to facilitate exchange and integration of data with other similar efforts as well as to facilitate human and machine based data mining. These personnel will be trained in the use of the latest pathway, complex, interaction, model organism and protein databases. They will act as a liason between researchers and databases to facilitate retrieval of information AND entry of curated information by local researchers.
Biomolecular interaction data consists of the set of experimentally verified interactions that occur between proteins, DNA’s, RNA’s, small molecules or complexes involving any of these molecular types. These data, along with associated reactions and state changes form the basis of biological pathways. As such, interaction and pathway data define the biological function of their participant molecules. The resulting network of interactions and pathways between molecules forms a map of living systems that may be searched and computed on. The resource is a broadly applicable to all molecular and medical life sciences.
Presently these data are collected by several small groups around the world. It is a labour intensive task that requires skill in reading research articles and knowledge of multiple standard data formats using large controlled vocabularies. Traditionally, these databases have had difficulties in securing long-term, stable funding since they compete with proposals for experimentalist research while they are essentially infrastructure projects. Despite the fact that these databases receive hundreds of citations per year, a survey of the major interaction databases indicates that they employ only a handful of full-time curators. This number is insufficient to keep up with the rate of research publications let alone the backlog of uncurated research articles. This infrastructure call represents a unique opportunity for Norway to establish a prototype position in this area that could be replicated across universities. The cost is fractional compared to the funds expended by universities on biomedical journal subscriptions. The payoff is a powerful dataset that may be data mined by humans and machine algorithms.
We will solicit letters of support from international interaction databases supporting our efforts in this area. We will also solicit national and international research groups to propose biomedical areas requiring curation. Finally, we will survey universities worlwide to assess their support of such a service. Newly curated data will be used to give context to high and low throughput proteomics and sequencing projects as well as provide tools for genome wide analysis studies related to human disease, cancer and personalized medicine. Graphical algorithms acting on large scale interaction and pathway maps have broad utility that includes (but is not limited to) identification of biological roles of proteins, identification of disease genes and selection of drug targets. The efficacy of these algorithms is dependent on the quality of the underlying data. Presently, high-quality, human curated data is dwarfed by less reliable data from high-throughput interactomics studies. The interpretation of these high-throughput studies themselves are benefited by the presence of human-curated data.
The proposed project will have high visibility and high impact. Data will be made freely available in internationally recognized formats (such as the HUPO PSI-MI standard) under a Creative Commons License. Data will be available via bulk-download, web-interface and at least one internationally recognized graphical viewer (http://cytoscape.org). Data will be integrated and exchanged with other similar database efforts to facilitate search and analysis. Integration will be accomplished using a system recently developed in the principal investigator’s research group (http://irefindex.uio.no). This same system will be used to monitor and ensure accepted curation practices. We will contribute to the maintenance and expansion of data exchange formats and controlled vocabularies. We will adhere to and develop curation practices set out by the International Molecular Exchange Consortium (http://imex.sourceforge.net/). Existing curation and database systems for handling data are already available from other IMEx groups and these will be installed, used and built upon.
This initiative is a proposed infrastructure project at the University of Oslo in Norway where this position type would be prototyped. The initiative is led by Ian Donaldson at the Biotechnology Centre of Oslo. The above survey is an attempt to assess support for the proposal at the University of Oslo and at Universities around the world where this project could be replicated.
How to add your university to this survey
We have sent query emails to lists of biology researchers at a number of universities. If you would like to do the same for your university, you can use the following message. Simply copy and paste. Results of the survey will be posted on this site.
Subject line: would you like to meet with a biolibrarian?
We are proposing the creation of a new infrastructure position at university libraries around the world. The position is called a “Biolibrarian”.
A Biolibrarian is trained in the use of biological databases that include biological pathway, complex and interaction databases.
It is envisioned that molecular biologists could meet with a Biolibrarian in the same way that they meet with and use the services of a librarian. The Biolibrarian could help molecular biologist researchers to locate pathways, complexes and interactions that their molecules of interest are involved in. The Biolibrarian would help the biologist to access, use and interpret data from curated molecular databases (including pathway, complex, interaction, model organism and protein databases).
Finally, Biolibrarians could help researchers enter verified information from full-text articles into curated databases where it would be available to researchers around the world that were querying for information on these same molecules.
Would you support such a service at your local university library? Do you have comments on this proposal? Please take our six question survey to enter your opinion.
http://www.surveymonkey.com/s.aspx?sm=LubRkhzbX9a7e8aPGKGevQ_3d_3d
You can read more about the proposal here: http://irefindex.uio.no/wiki/The_Biolibrarian_Proposal
About the author and contact information
This proposal was written by Ian Donaldson. I was a lead bioinformatics developer and research scientist for the Biomolecular Interaction Network Database (BIND) between 2002 and 2005. This effort employed close to 30 curators. I was involved in many aspects of this project (including curation and data standard development) since the project’s inception in 1999.
This proposal is an attempt to secure long term funding and support for my own research efforts (iRefIndex) and the protein-interaction databases that were included in this initial study. However, in my own experience, I believe that funding of databases on a per research group basis is an unsustainable solution. Therefore, I have caste this proposal in the light of a more global and inclusive solution that will support mature, biological databases in general. Again, based on my own experience, I believe that there is a clear need for an intermediary between biological databases and local researchers: this is the basis of the Biolibrarian proposal. I believe that a global-local network of such specialists should eventually work at arms length from any research group (including my own) and be responsible only to their host universities as an infrastructure resource. This proposal suggests prototying such a group to demonstrate its possibilities and potentials. I welcome feedback, comments and criticisms of these ideas.
Ian Donaldson
Ian Donaldson, Ph.D.
Biotechnology Centre of Oslo, University of Oslo
Visiting addr: Gaustadalléen 21, 0349 Oslo
Postal addr: P.O. Box 1125 Blindern, 0317, Oslo
Phone: +47 99 11 51 49
Fax: +47 22 84 05 01
Email: ian.donaldson@biotek.uio.no
Skype: ian.oslo
Related sites
Big data: the future of biocuration. See full article and PMID 18769432.